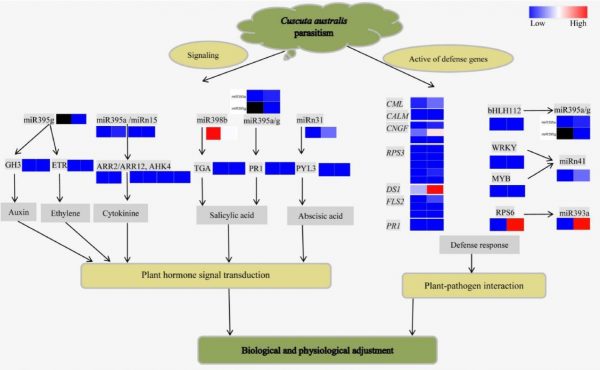
Cuscuta australis R.Br. is a stem parasite that obtains water and nutrients from host plants, with which it also exchanges proteins and mRNAs with host plants. Recently, a study found that novel miRNAs accumulated during C. campestris Yunk. parasitism of Arabidopsis thaliana L. However, the fate of miRNAs transferred from parasite to host plants as well as their targets has remained unexamined. To understand the miRNA–mRNA interaction network of host Trifolium repens L. in response to C. australis parasitism, integrated mRNA, small RNA, and degradome sequencing data were analyzed. In total, 60,824 unigenes were identified by RNA-seq, of which 1,601 were differentially expressed genes (DEGs). A total of 71 known miRNAs belonging to 38 miRNA families and 65 novel miRNAs were identified from small RNA sequencing data and 8,012 target genes were predicted. Of these target genes, 129 miRNA–mRNA pairs were identified via degradome analyzes. There were also eight differentially expressed miRNAs (DEMs). DEG and DEM correlation analyzes revealed 23 pairs of differentially expressed miRNA–mRNA pairs responsive to C. australis parasitism. The most enriched pathways were plant hormone signal transduction, ribosome and plant–pathogen interaction pathways. The miRNA–mRNA pairs in host T. repens, including trr-miR393a–RPS6, trr-miR398b–TGA, trr-miR395a/g–bHLH112, trr-miR395g–GH3, and trr-miRn41–WRKY/MYB, likely play important roles in interactions between host T. repens and parasitic C. australis. These results provide valuable information on the mechanisms of parasite–host plant interactions. (Scientia Horticulturae, 2021, 289: 110458)